Abstract
Background Artificial intelligence (AI) tremendously influences our daily lives and the medical field, changing the scope of medicine. One of the fields where AI, and, in particular, predictive modeling, holds great promise is spinal oncology. An accurate patient prognosis is essential to determine the optimal treatment strategy for patients with spinal metastases. Multiple studies demonstrated that the physician’s survival predictions are inaccurate, which resulted in the development of numerous predictive models. However, difficulties arise when trying to interpret these models and, more importantly, assess their quality.
Objective To provide an overview of all stages and challenges in developing predictive models using the Skeletal Oncology Research Group machine learning algorithms as an example.
Methods A narrative review of all relevant articles known to the authors was conducted.
Results Building a predictive model consists of 6 stages: preparation, development, internal validation, presentation, external validation, and implementation. During validation, the following measures are essential to assess the model’s performance: calibration, discrimination, decision curve analysis, and the Brier score. The structured methodology in developing, validating, and reporting the model is vital when building predictive models. Two principal guidelines are the transparent reporting of a multivariable prediction model for individual prognosis or diagnosis checklist and the prediction model risk of bias assessment. To date, many predictive modeling studies lack the right validation measures or improperly report their methodology.
Conclusions A new health care age is being ushered in by the rapid advancement of AI and its applications in spinal oncology. A myriad of predictive models are being developed; however, the subsequent stages, quality of validation, transparent reporting, and implementation still need improvement.
Clinical Relevance Given the rapid rise and use of AI prediction models in patient care, it is valuable to know how to assess their quality and to understand how these models influence clinical practice. This article provides guidance on how to approach this.
Level of Evidence 4.
- artificial intelligence
- machine learning
- orthopedic surgery
- prediction tools
- clinical decision support
- spinal oncology
Introduction
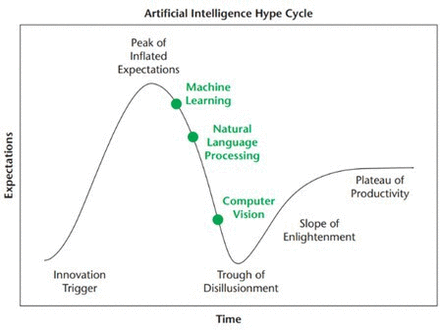
Artificial intelligence (AI) tremendously influences not only our daily lives but also the medical field, changing the scope of medicine. Improvements in computational power, along with AI-based software platforms, and the availability of more extensive electronic data, have enabled the development of many different applications, such as machine learning (ML)–derived clinical decision support tools, deep learning-based computer vision, and natural language processing.1 Oosterhoff et al suggested in 2020 that we have reached the peak of inflated expectations in medical AI along with Gartner’s hype cycle (Figure 1).2 Although the promise of AI remains strong, where an individual stands on the hype cycle would depend on their experience and understanding of AI. Individuals new in this field can still be at the peak of inflated expectations, while more experienced individuals might be toiling through the trough of disillusionment as challenges in implementing AI applications are becoming more apparent. The purpose of the present article is to provide a narrative review of AI and predictive modeling in spinal oncology and discuss the potential and limitations of the technology. We present no unpublished data and reference to data from previously published studies.
Gartner’s hype cycle. Source: Reprinted with permission from Oosterhoff JHF, Doornberg JN. Artificial intelligence in orthopaedics: false hope or not? A narrative review along the line of Gartner’s hype cycle. EFORT Open Rev. 2020;5(10):593–603. © 2020 Oosterhoff and Doornerg.
Spinal Oncology
One of the fields where AI, and, in particular, predictive modeling, has made significant advances is spinal oncology. The spine is the most common location of metastatic cancer disease,3–5 and 30% to 90% of patients who die of cancer have spinal metastasis in cadaver studies.6–9 Up to 50% of spinal metastasis require treatment, and 5% to 10% need surgical management.8,10,11 Moreover, cancer survival rates are increasing due to earlier detection and improved treatment, and the prevalence of spinal metastasis will also likely increase.12 In 2005, the landmark article of Patchell et al13 showed that surgical intervention is efficacious in treating metastatic spinal tumors. Following this, together with the emergence of a myriad of treatments, including personalized systemic therapy and targeted therapy, a systemic decision framework for treating spinal metastases was necessary.14 In 2015, the neurologic, oncologic, mechanical, and systemic decision framework was developed to determine the optimal therapy for patients with spinal metastases.15 This framework enabled physicians to apply a systematic approach to treating spinal metastases, resulting in an increased surgery rate.16 However, spinal surgery is not without risk; surgery complications are a significant source of comorbidity and include wound infections, neurologic impairment, venous thromboembolism, instrumentation failure, and pain.17–20 Moreover, patients with metastatic spinal disease generally have multiple medical comorbidities and are immunocompromised due to immune suppression.21 Therefore, treatment goals focus on whether patients will likely recover from the indicated procedure.22 The appropriate use of surgery for metastatic spinal disease is dependent on the expected risk of surgery and the expected benefit. Accurate expectations for risk and benefit would be valuable to empower informed choice for physicians and patients.
The Emergence of Prediction Tools
Multiple studies have shown that physicians’ clinical predictions of the life expectancy of cancer patients are inaccurate.23,24 In 2005, Nathan et al showed that a better means of prognostication was needed.25 Consequently, numerous new scoring systems and prognostic calculators were developed.26–36 Unfortunately, many did not meet the required accuracy, performed inconsistently, or lacked personalized predictions.26,37 Thirteen survival prediction scores exist, including PATHFx,38 Skeletal Oncology Research Group ML algorithms (SORG-MLA),33 Bollen Classification,39 modified Bauer score,34 and van der Linden40 (Supplement 1).27,41–47 Of these prediction scores, SORG-MLA and PATHFx are the only 2 ML algorithms. Over the past years, SORG-MLA demonstrated its clinical value and promise over other prediction scores such as nomograms or regression models. However, important questions regarding the use of AI in predictive models remain, including the following: (1) How do we interpret prognostic AI models such as SORG-MLA? (2) How do we assess their quality? and (3) How will these models influence clinical practice?
Figure S1.
Figure S2.
Development, Validation, and Implementation of Prediction Models
Why Machine Learning?
Statistical models have been widely used to formalize the understanding of data, but since data size and variable inputs increased, these models have become more complex. Fortunately, ML models have become more powerful due to an increase in computational power. According to Bzdok et al,48 “statistics draws population inference from a sample, and ML finds generalizable predictive patterns.” In principle, many methods from statistics and ML can be used for both prediction and inference. However, statistical methods have a long-standing focus on inference, achieved through creating and fitting a project-specific probability model. In contrast, ML concentrates on prediction with general purpose learning algorithms to find patterns in often rich and unwieldy data.49,50 They are particularly helpful when dealing with “wide data,” where the number of input variables exceeds the number of subjects. Thus, where statistical models are generally hypothesis-driven, ML is more exploratory in identifying correlations, and the pattern of correlation is not a causal relationship. This may be recognized as a limitation of ML. However, with the possession of large patient data sets due to electronic health care systems, ML provides the opportunity to find patterns and determine values predictive of the output requested. Therefore, ML offers a more accurate solution for developing prediction models, such as the survival probability of patients with metastatic spine disease, which is complicated and requires multiple aspects to be considered.
Steps in Building Predictive Models
Structured methodology in the development and validation of an ML model is of great importance and is best executed along the ABCD steps of Steyerberg et al.51 Additionally, 2 important guidelines are important to adhere to: the transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD)52 checklist, essential for transparent reporting of a prediction model study, and the prediction model risk of bias assessment (PROBAST),53 a tool for assessing the risk of bias and applicability of prediction model studies. With the SORG-MLA for 1-year survival, developed and validated multiple times within our research team, as an example, we will go through the steps of model preparation, development, validation, presentation, and implementation (Figures 2–4).54–57
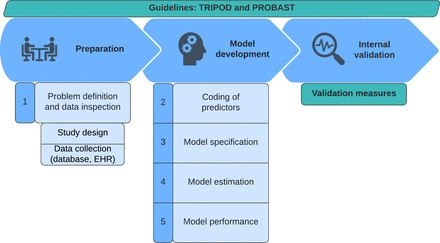
The first 3 stages in model development: preparation, development, and internal validation. TRIPOD, transparent reporting of a multivariable prediction model for individual prognosis or diagnosis; PROBAST, prediction model risk of bias assessment tool; EHR, electronic health record.
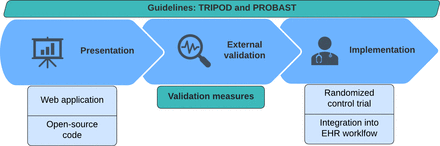
The last 3 stages in model development: presentation, external validation, and implementation. TRIPOD, transparent reporting of a multivariable prediction model for individual prognosis or diagnosis; PROBAST, prediction model risk of bias assessment tool; EHR, electronic health record.
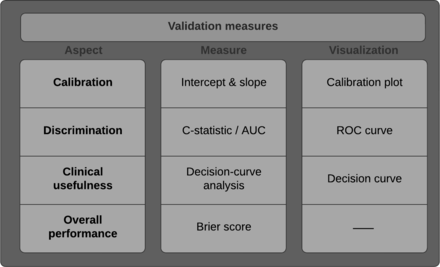
Overview of validation measures. C-statistic, concordance statistic; AUC, area under the curve; ROC, receiver operating characteristic.
The first step is the consideration of the research question and initial data inspection. For the development of the SORG-MLA, the objective was to find predictive variables and develop a predictive algorithm for survival of metastatic spinal disease at intermediate (90-day) and long-term (1-year) time points. Based on the expert knowledge and previous literature, we chose a framework of input variables to consider. Patients were included when they were older than 18 years, had a diagnosis of metastatic spinal disease, and had an initial surgical procedure performed between 1 January 2000, and 31 December 2016. Missing data were imputed with the missForest multiple imputation method, which is currently considered one of the superior imputation methods. Baseline data collection was retrospective, and the definitions of all input variables, generally referred to as predictors, were carefully documented.
The second step is the coding of the predictors. Categorical and continuous predictor variables can be coded in different ways. At the start of model development, coding the variables in a detailed way is preferred so that in a later phase, when relative effects of predictors are known, a user-friendly variable format may be used. For example, when coding the variable of primary tumor histology, we might see that coding the variable in 3 groups according to primary tumor instead of coding them all separately would result in similar performance, making the model simpler to use.
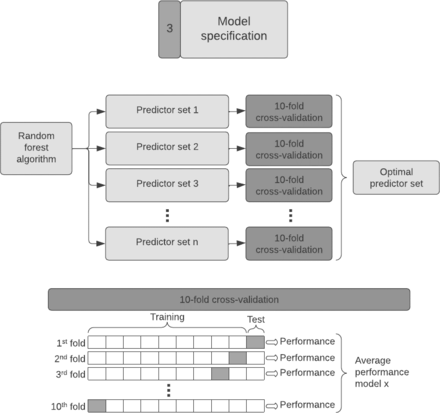
The third step is the model specification, where we choose the predictors for inclusion in the prediction model (Figure 5). For SORG-MLA, we used random forest algorithms with 10-fold cross-validation, which enabled us to find the optimal subset of predictors while keeping the variance of the model performance low and avoiding overfitting.
Model specification. With a random forest algorithm, we created many different predictor sets (sets with different input variables) which we tested with 10-fold cross-validation to find the optimal set of predictors. This technique fits the model 10 times, with each fit being performed on a training set of a different 90% of the data with the remaining 10% as a holdout set for validation. Each fit produces a performance metric, and the average of all these fits results in the average performance of a predictor set.
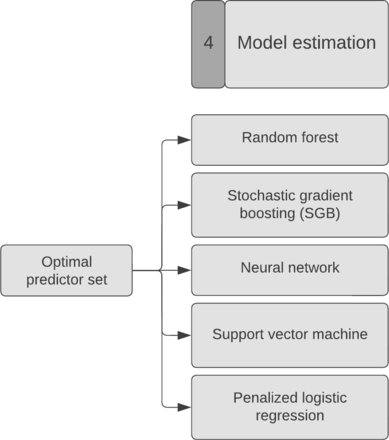
The fourth step is the model estimation: choosing the right ML model (Figure 6). For SORG-MLA, we used 5 different models based on a previous study’s method.58 The data were then divided into a training set (80%) and a holdout validation set (20%). The training set is used to train the models, and the validation set is used to internally validate the model. An independent validation set is essential to test the models on unseen data.
Model estimation. For SORG-MLA, we used 5 different models: random forests, stochastic gradient boosting, neural network, support vector machine, and penalized logistic regression. SORG-MLA, the Skeletal Oncology Research Group machine learning algorithms.
The fifth and sixth steps are the validation and evaluation of model performance, where we determine the quality and performance of the algorithms and alter the algorithm if necessary. Evaluation and validation are ideally performed along the ABCD steps; these will be discussed in the next section.
The seventh, and final, step is the model presentation such that it best addresses the clinical needs. We presented SORG-MLA as an open access web-based application to facilitate accessibility (see https://sorg-apps.shinyapps.io/spinemetssurvival/). However, ultimately, integration into decision aids and electronic patient records will best support clinical decision-making.59
Validation Methods
Model validation is the process by which predictions are compared with independent real-world observations to judge quantitative and qualitative properties of the model. There are 4 important measures based on the “ABCD” steps of Steyerberg et al,51 which together provide an accurate and well-established validation and evaluation: calibration, discrimination, decision curve analysis, and the Brier score.38,57,60,61
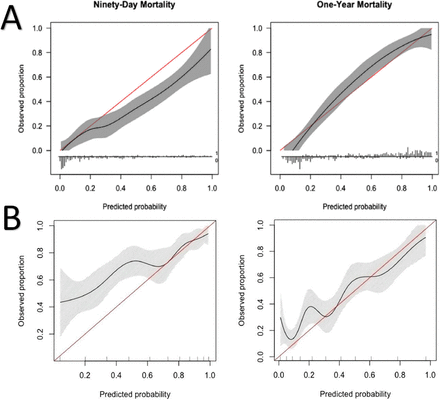
Calibration (A and B) refers to the agreement between observed end points and predictions and answers the question: Is the model as reliable when it predicts a 10% probability as when it predicts a 70% probability of mortality?62 It can be best assessed graphically in a calibration plot with survival predictions on the x-axis and real-world observations on the y-axis. Perfect calibration of a model should have a straight line, described with an intercept of 0 and a slope of 1. Imperfect calibration can be observed by deviation from this ideal straight line (Figure 3). This calibration plot helps visualize whether models overestimate or underestimate the outcome. The SORG-MLA achieved an intercept of 0.07 and a slope of 1.26 (Figure 7), showing a near perfect intercept and a slightly higher slope, indicating that there are individuals or subgroups in whom calibration is suboptimal and survival is overestimated.63
Calibration: calibration plot of SORG-MLA predicting 90-d and 1-y mortality at (A) internal validation and (B) external validation (Taiwan). Comparing these plots demonstrates that SORG-MLA performs differently in other populations, highlighting the importance of external validation. SORG-MLA, the Skeletal Oncology Research Group machine learning algorithms. Source: Reprinted from with permission from The Spine Journal, Vol 21, Yang J-J, Chen C-W, Fourman MS, et al, International external validation of the SORG machine learning algorithms for predicting 90-day and one-year survival of patients with spine metastases using a Taiwanese cohort, 1670-1678, Copyright 2021, with permission from Elsevier.56
Discrimination (C) refers to the ability of the model to distinguish the end points, that is, whether a patient is dead or alive at the specified time point. The measure is quantified by the area under the curve of the receiver operating characteristic curve, which represents the probability that the model will be able to differentiate between patients who survived and those who died. Interpretation of this curve can be simplified: 0.51 to 0.69 poor, 0.70 to 0.79 fair, 0.80 to 0.89 good, 0.90 to 0.99 excellent. The SORG-MLA achieved an area under the curve of 0.89.63
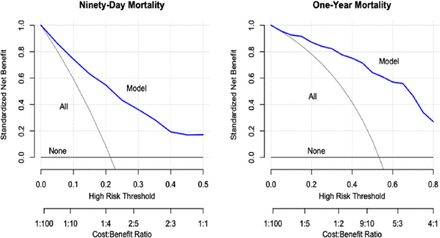
Even though calibration, discrimination, and the Brier score are essential, these measures do not assess the clinical usefulness or the ability to make better clinical decisions with the model than without. To determine the impact of these models on clinical decisions, it is essential to perform a decision analysis (D). Even though this type of analysis has been around for a significant amount of time, it only recently started gaining popularity as a necessary tool in prediction models.62,64 Decision curve analysis examines the net benefit of decisions made based on the model predictions. Changing management for all patients and changing management for no patients are the 2 default strategies for decisions without prediction models. Decision curves show whether the clinical prediction model used for management changes offers a greater net benefit than the 2 default strategies. The SORG-MLA showed greater standardized net benefit at all predicted probabilities relative to management decision change based on treating all patients or no patients (Figure 8).57
Decision curve analysis: decision curve of SORG-MLA predicting 90-d and 1-y mortality at external validation. SORG-MLA, the Skeletal Oncology Research Group machine learning algorithms. Source: Reprinted from The Spine Journal, Vol 21, Shah AA, Karhade AV, Park HY, et al, Updated external validation of the SORG machine learning algorithms for prediction of ninety-day and one-year mortality after surgery for spinal metastasis, 1679–1686, Copyright 2021, with permission from Elsevier.57
Another important measure, although not recorded in the ABCD steps, is the Brier score: a summary measure that formalizes the performance of predictions. The so-called “null model” of the Brier score corresponds to the scenario where every patient is predicted to have a risk equal to the prevalence of mortality in the whole disease population. The Brier score calculates the error between the prediction and observed outcome for each patient and compares it to the null model. Ideally, zero error between the predictions and outcomes is preferred, resulting in a perfect Brier score of 0. The SORG-MLA achieved a Brier score of 0.13, whereas the null model had a Brier score of 0.25.63
Validation of the model can only be adequately assessed when all measures are performed. For example, a model can have excellent discrimination but very poor calibration. Or, a model could have good discrimination and calibration but worse standardized net benefit compared with default changes in management, resulting in a model that harms clinical decision-making. Therefore, assessing and reporting every validation measure mentioned above are essential.
Internal and External Validation
Assessing model validation is executed at 2 stages: internal validation at the end of model development and external validation when the model is already presented. The difference is that internal validation is performed at the institute that develops the model, whereas external validation is done at multiple (different) institutions, assessing the model’s generalizability to different patient populations. When validating a prediction model, it is important to not only assess the measures mentioned before but also assess whether the model has been developed correctly. To facilitate this, transparent and complete reporting of the development and validation of a model are required to allow the reader to critically assess the presence of bias, facilitate study replication, and correctly interpret results.65 External validation of SORG-MLA has been done extensively in the United States and multiple international patient populations (Table).54–57 However, the overall survival of patients with spinal metastases is improving and will hopefully keep improving due to improved treatments and clinical decision-making.12 This may result in lower performance of the model in the future. Therefore, it is vital to continuously monitor and validate the performance of ML models so that clinicians and data scientists can identify and assess performance deviations as soon as possible and recalibrate or update models if necessary.
External validations of SORG-MLA predicting 90-d and 1-y mortality.
Implementation
Once external validation has been successful, the next step is implementing the model into clinical practice. An essential factor for integrating a model into clinical practice is ensuring clinicians’ trust and accurately interpreting the model.66 To earn this trust, transparent reporting of the model’s development, internal, and external validation is essential. Next, we must assess the real-world performance of the model on operational data, thus validating the algorithm on a prospective cohort by comparing the model performance with a surgeon with or without the model. Consequently, the performance of the developed model is ideally assessed with randomized control trials. Guidelines such as CONSORT-AI (Consolidated Standards of Reporting Trials - Artificial Intelligence) and SPIRIT-AI (Standard Protocol Items: Recommendations for Interventional Trials - Artificial Intelligence) have been developed to assist in the application for these trials.
To facilitate easy access to SORG-MLA, we presented the model as an open access web application. However, a real-time outcome calculator based on the developed ML algorithm and routinely collected data is best established, validated, and integrated within the electronic health record (EHR) systems.59 This has implications for patient privacy and creates obstacles for implementation.67 For SORG-MLA, we are currently performing an international, multicenter prospective study to evaluate the survival predictions of surgeons with or without the model. Consequently, if the study shows survival predictions improve significantly, implementation into EHR will follow.
Recommendations and Challenges
Despite the potential increased benefit of predictive models, there are restrictions and risks associated with ML models. As we have gone through all aspects of model processing, we will highlight several challenges in each stage.
Preparation
The quality of the data from which prediction models are produced determines the quality of those models.68 Even if the amount of data is large, data inaccuracy and missing data still pose serious problems when EHR data are used and may impact prognostic factors, treatment exposures, and outcome estimation.69 Because existing ML models are created using small, retrospective cohorts or registries, they frequently lack generalizability. This is particularly problematic in ML algorithms as they tend to amplify the biases and confounds already present in a dataset. Therefore, the PROBAST bias tool is so important. To increase the available data, many institutions are setting up multicenter or international databases or registries. However, these may be constrained by varied terminology affecting data labeling.
Considering the cost and time needed to utilize predictive modeling, spine surgeons, oncologists, and researchers should balance the upfront investment of time and money required to develop and validate predictive models.70 Predictive ML models can assist clinicians, but if there is no apparent need for more accurate predictions or if simple statistical models suffice, developing these models would not necessarily be advantageous.
Development and Validation
Even though there has been a massive increase in the volume of predictive models, quality and transparent reporting were not performed consistently. Quality of reporting refers to the application and reporting of the established validation measures. Unfortunately, of 18 studies externally validating 10 different ML prediction models in orthopedic surgery, only 39% reported calibration and 50% reported decision curve analysis.71 Transparent reporting refers to whether an article mentions all required items in development and validation recommended by the TRIPOD checklist and PROBAST tool. A recent study by Groot et al65 showed that in ML studies in orthopedics, adherence to the TRIPOD guidelines and PROBAST bias tool was limited. They reviewed 59 ML prediction studies published in orthopedic surgery, of which 18 (31%) were in the spine. The overall completeness for the TRIPOD checklist was 53%, and the overall risk of bias was low in 44%, high in 41%, and unclear in 15%.65
These results show that many studies incompletely reported their methods and performance measures. This, together with the fact that the relative novelty of this technique is viewed skeptically, makes it harder for clinicians to rely on predictive models. Thus, to enable trust and facilitate implementation, adherence to the guidelines and transparent reporting of these steps are essential. Consequently, TRIPOD-AI and PROBAST-AI were recently proposed for explicit use in AI to further aid in directing the future of this field.72
Even so, the aforementioned performance evaluations might not be sufficient to identify harmful or uninformative algorithms.69 Moreover, recent research has demonstrated that models created using retrospective data may be biased against racial minorities.73 Last, many AI algorithms are referred to as black boxes: we are unaware of the operations between input and output. Thus, fully interpreting the models becomes difficult. For this reason, the website of SORG-MLA contains explanations for which predictors contradict or support the model, allowing clinicians to interpret and explain the predicted mortality.
Implementation
Aside from challenges in the development and validation, more challenges arise when implementing ML models in clinical practice. As mentioned before, randomized prospective trials are essential to compare the accuracy of the survival prediction of a surgeon with or without the model. However, very few trials have been performed for predictive models in medicine and, to our knowledge, none to date in orthopedics or spine.61,69,74 Additionally, ethical, legal, political, and administrative barriers must be overcome. Ethical concerns include liability in cases of medical error, doctors’ understanding of how these models produce predictions, and patients’ understanding and control of how these models are used in their care.75 Moreover, issues of privacy, security, and management of patient data are important to consider.
Conclusion
A new health care age is being ushered in by the rapid advancement of AI and its applications in spinal oncology. A myriad of new models are being developed, but the subsequent stages, quality of validation, transparent reporting, and implementation still need improvement. Moreover, we must acknowledge that these models are not a single means to an end. When interpreting these algorithms, we must always consider the context of the clinical question regarding the patient. It will be vital as we advance to regularly scan for potential dangers and ensure that patient benefit and safety continue to come first.
Footnotes
Funding The authors received no financial support for the research, authorship, and/or publication of this article.
Declaration of Conflicting Interests The authors report no conflicts of interest in this work.
Disclosures Each author certifies that he or she has no commercial associations (eg, consultancies, stock ownership, equity interest, patent/licensing arrangements, etc) that might pose a conflict of interest in connection with the submitted article. Investigation performed at Massachusetts General Hospital, Boston, USA.
- This manuscript is generously published free of charge by ISASS, the International Society for the Advancement of Spine Surgery. Copyright © 2023 ISASS. To see more or order reprints or permissions, see http://ijssurgery.com.